REDDI: A Riemannian Ensemble Learning Framework for Interpretable Differential Diagnosis of Neurodegenerative Diseases
Abstract
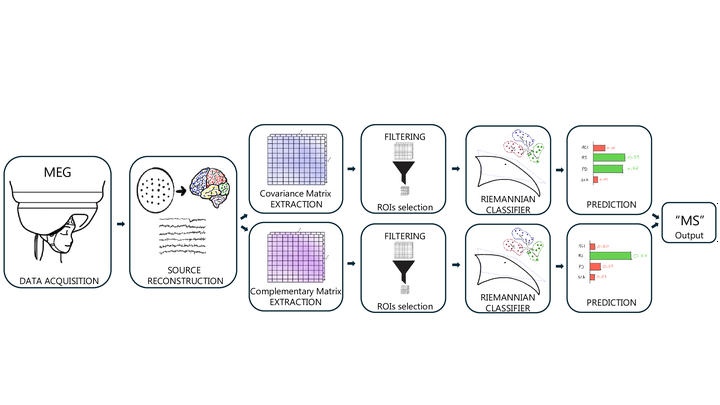
Neurodegenerative diseases such as Mild Cognitive Impairment (MCI), Multiple Sclerosis (MS), Parkinson s Disease (PD), and Amyotrophic Lateral Sclerosis (ALS) are becoming more prevalent. Each of these diseases, despite its specific pathophysiological mechanisms, leads to widespread reorganization of brain activity. However, the corresponding neurophysiological signatures of these changes have been elusive. As a consequence, to date, it is not possible to effectively distinguish these diseases from neurophysiological data alone. This work uses Magnetoencephalography (MEG) resting-state data, combined with interpretable machine learning techniques, to support differential diagnosis. We expand on previous work and design a Riemannian geometry-based classification pipeline. The pipeline is fed with typical connectivity metrics, such as covariance or correlation matrices. To maintain interpretability while reducing feature dimensionality, we introduce a classifier-independent feature selection procedure that uses effect sizes derived from the Kruskal-Wallis test. The ensemble classification pipeline, called REDDI, achieved a mean balanced accuracy of 0.81 (+/-0.04) across five folds, representing a 13% improvement over the state-of-the-art, while remaining clinically transparent. As such, our approach achieves reliable, interpretable, data-driven, operator-independent decision-support tools in Neurology.